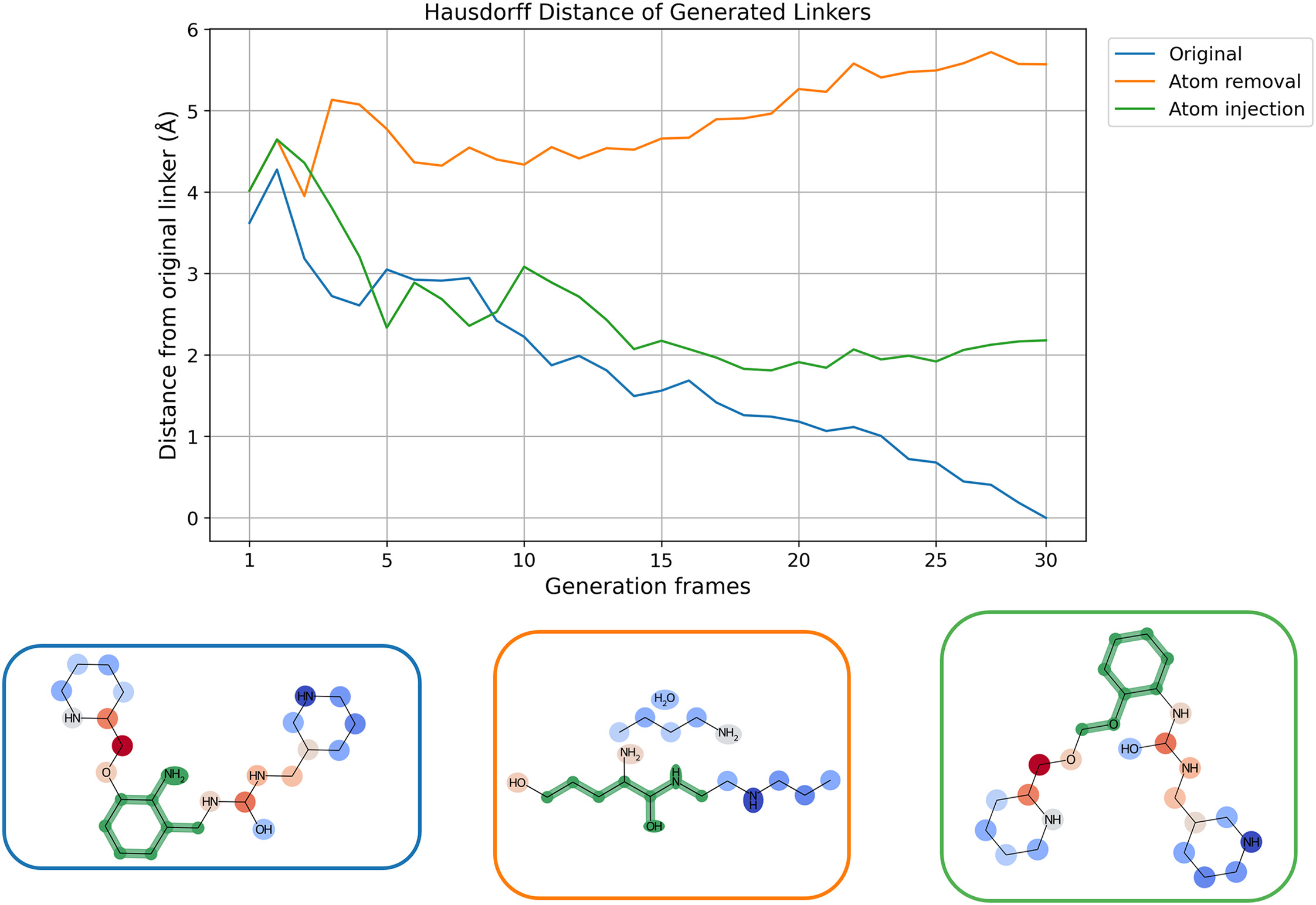
Diffusion models are increasingly being used in drug discovery. However, the mechanisms at work within these models are still only partially understood. A recent study, published in Cell Reports Physical Science, by Dr. Andrea Mastropietro and Prof. Dr. Jürgen Bajorath from the University of Bonn and the Lamarr Institute for Machine Learning and Artificial Intelligence now investigates how these AI models actually work and shows that they utilize chemical relationships differently than has often been assumed.
Diffusion models are a form of generative AI. They generate new data by gradually removing so-called noise from existing examples. Noise refers to random variations that overlay an original pattern. What was initially used for images and videos is now also being applied to molecular design tasks. The study focuses on linker design. Linkers are molecular structures that connect separate parts of a molecule and significantly influence its properties, such as how well an active ingredient binds to its target.
How the model assembles molecules
To analyze the generation process, the researchers developed a method called DiffSHAPer. It is based on the concept of Shapley values from explainable AI. Shapley values originate from game theory and describe the contribution individual elements make to an overall outcome. In this context, they make it possible to quantify how strongly individual atoms of a molecular fragment influence the generation of a linker. This reveals which parts of a molecule have a particularly strong influence on the formation of a linker.
The results show that, when generating chemically valid linkers, the model under investigation primarily relies primarily on spatial distances between atoms. The researchers found no evidence that it systematically uses generalizable chemical rules or functional relationships in this process. Instead, the generation appears to be significantly shaped by recurring statistical patterns in the training data.
What this means for research and practice
For drug discovery, this means that while diffusion models do indeed yield formally correct molecular structures, their functional properties cannot be directly inferred from them. Whether these structures possess the desired properties, such as stability or the ability to bind specifically to a biological target, is therefore not automatically guaranteed. A linker based primarily on geometric criteria may be functionally unsuitable. The results thus touch on a central question in current AI research: To what extent do generative models capture underlying chemical relationships, or do they primarily reflect statistical correlations? This distinction is particularly crucial in scientific fields such as drug development, as misconceptions about a model’s internal mechanisms can have direct implications for research outcomes. This work aligns with the Lamarr Institute’s research on trustworthy and explainable AI. The focus is on analyzing complex models whose results must be verifiable and scientifically interpretable. The study provides an example of how generative methods can be examined in detail and enables the targeted further development of such models.